- Played:94,357
Do you like this game?
This game on a smartphone:
Embed Code:
Share with Friends:
The Tower of Hanoi is a puzzle game invented in 19th century by the French mathematician Eduard Lucas. This guy got inspired by the legend about priests in Hindi castle. Those priests got 3 bars, one of them had 64 golden disks. Each bar was slightly larger than the previous one. As the legend says, they had to move the tower from one bar to another one following the two conditions. First of all, they could only move one disk at a time. Secondly, they were not allowed to put a larger disk on a smaller one. According to the legend, when they completed the work, the castle would turn into dust, and the world would vanish. We calculated that 264-1=18 446 744 073 709 551 615 steps are needed to solve the puzzle with 64 disks. If 1 second is spent to complete each step, the tower will be moved during 586 billion years. The Earth is about 4 and a half billion years old now.
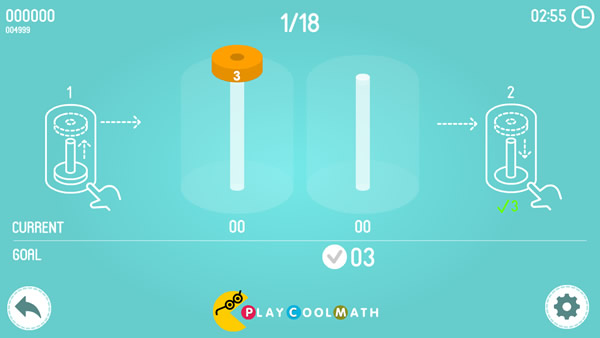
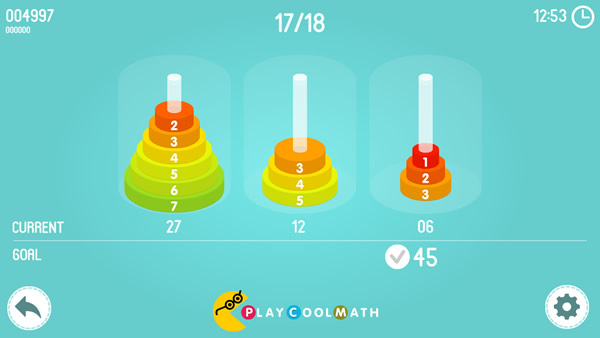
Based on this puzzle, we created a game called Math Tower of Hanoi. The game contains of 18 levels - from simple ones to complex ones. The smallest disk has number 1 on it, the next disk is larger and has number 2 and so on up to the largest disk that has number 8, accordingly. The main goal of this game is to move the disks so that the amount of the digits on all the disks in the bars would match the target. The fewer steps you make during the game, the more points you will get for the puzzle. You are allowed to move only one disk at a time. You cannot put a larger disk on a smaller one. You will get other math tasks between the levels. Play and improve your skills in mental counting.
Ready to try?